If you already understand Simple Linear Regreession, then we can make things a little more complicated. Multivariate Linear Regression considers inputs with multiples features. This will be help us to develop a dense layer for the next part.

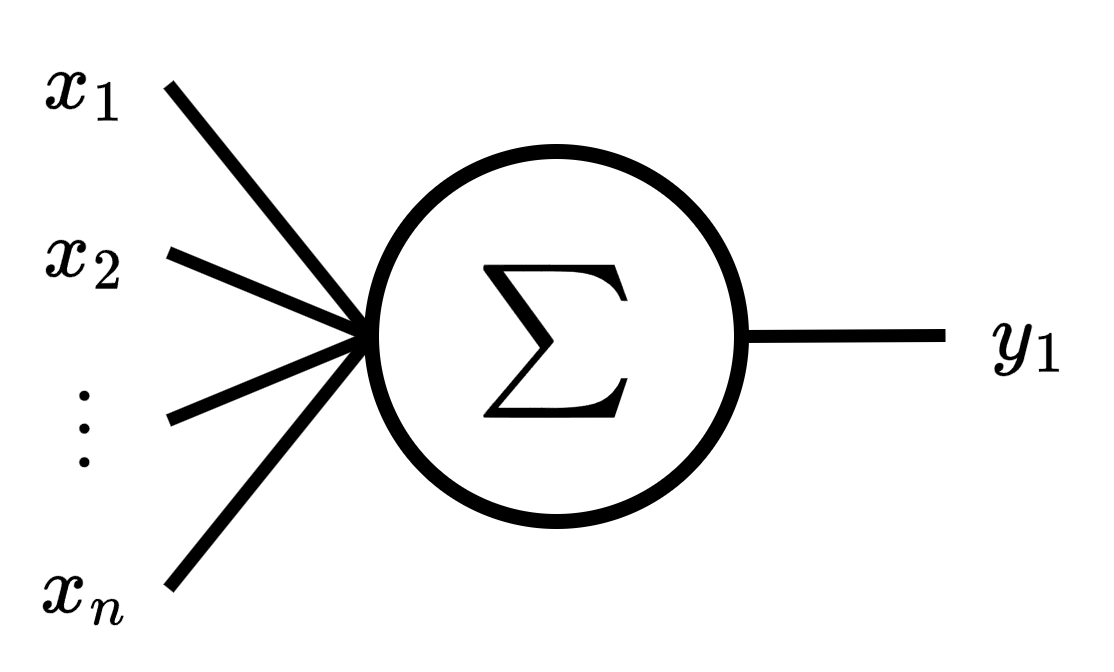
The goal of multivariate linear regression is similar to simple linear regression, estimate by a linear approximation
Note that input data is now a matrix.
Purpose of this Notebook:
Create a dataset for multivariate linear regression task
Create our own Perceptron class from scratch
Calculate the gradient descent from scratch
Train our Perceptron
Compare our Perceptron to the one prebuilt by PyTorch
Setup¶
print('Start package installation...')Start package installation...
%%capture
%pip install torch
%pip install scikit-learnprint('Packages installed successfully!')Packages installed successfully!
import torch
from torch import nn
from platform import python_version
python_version(), torch.__version__('3.12.12', '2.9.0+cu128')device = 'cpu'
if torch.cuda.is_available():
device = 'cuda'
device'cpu'torch.set_default_dtype(torch.float64)def add_to_class(Class):
"""Register functions as methods in created class."""
def wrapper(obj): setattr(Class, obj.__name__, obj)
return wrapperDataset¶
create dataset¶
The dataset is consists of the input data and the target data
The input data can be represented as a matrix
where is the number of samples, is the number of features, and .
The target data still without changes
from sklearn.datasets import make_regression
import random
M: int = 10_100 # number of samples
N: int = 4 # number of features
X, Y = make_regression(
n_samples=M,
n_features=N,
n_targets=1,
n_informative=N - 1, # let's add a features as a linear combination of others
bias=random.random(), # random true bias
noise=1
)
print(X.shape)
print(Y.shape)(10100, 4)
(10100,)
split dataset¶
X_train = torch.tensor(X[:100], device=device)
Y_train = torch.tensor(Y[:100], device=device)
X_train.shape, Y_train.shape(torch.Size([100, 4]), torch.Size([100]))X_valid = torch.tensor(X[100:], device=device)
Y_valid = torch.tensor(Y[100:], device=device)
X_valid.shape, Y_valid.shape(torch.Size([10000, 4]), torch.Size([10000]))delete raw dataset¶
del X
del YScratch multivariate perceptron¶
weight and bias¶
class MultiLinearRegression:
def __init__(self, n_features: int):
self.b = torch.randn(1, device=device)
self.w = torch.randn(n_features, device=device)
def copy_params(self, torch_layer: nn.modules.linear.Linear):
"""
Copy the parameters from a module.linear to this model.
Args:
torch_layer: Pytorch module from which to copy the parameters.
"""
self.b.copy_(torch_layer.bias.detach().clone())
self.w.copy_(torch_layer.weight[0,:].detach().clone())weighted sum¶
Remark: We can add a scalar to a vector due to broadcasting mechanism.
@add_to_class(MultiLinearRegression)
def predict(self, x: torch.Tensor) -> torch.Tensor:
"""
Predict the output for input x.
Args:
x: Input tensor of shape (n_samples, n_features).
Returns:
y_pred: Predicted output tensor of shape (n_samples,).
"""
return torch.matmul(x, self.w) + self.bMSE¶
MSE still without changes.
@add_to_class(MultiLinearRegression)
def mse_loss(self, y_true: torch.Tensor, y_pred: torch.Tensor):
"""
MSE loss function between target y_true and y_pred.
Args:
y_true: Target tensor of shape (n_samples,).
y_pred: Predicted tensor of shape (n_samples,).
Returns:
loss: MSE loss between predictions and true values.
"""
return ((y_pred - y_true)**2).mean().item()
@add_to_class(MultiLinearRegression)
def evaluate(self, x: torch.Tensor, y_true: torch.Tensor):
"""
Evaluate the model on input x and target y_true using MSE.
Args:
x: Input tensor of shape (n_samples, n_features).
y_true: Target tensor of shape (n_samples,).
Returns:
loss: MSE loss between predictions and true values.
"""
y_pred = self.predict(x)
return self.mse_loss(y_true, y_pred)gradients¶
Let’s follow the same strategy as before:
First, determine the derivatives to be computed
Then, ascertain the shape of each derivative
Finally, compute the derivatives
⭐️ We are using Einstein notation, that implies summation. For example
we will use Einstein notation for chain rule summation, for example
Derivative of MSE respect to bias
and derivative of MSE respect to weight
where the shape of each derivative is
MSE derivative¶
The vectorized form is
weighted sum derivative¶
respect to bias¶
for .
respect to weight¶
for , and .
Vectoring for all
Vectorizing for all
full chain rule¶
Derivative of MSE respect to bias
Vectorized form is
final gradients¶
parameters update¶
@add_to_class(MultiLinearRegression)
def update(self, x: torch.Tensor, y_true: torch.Tensor,
y_pred: torch.Tensor, lr: float):
"""
Update the model parameters.
Args:
x: Input tensor of shape (n_samples, n_features).
y_true: Target tensor of shape (n_samples,).
y_pred: Predicted output tensor of shape (n_samples,).
lr: Learning rate.
"""
delta = 2 * (y_pred - y_true) / len(y_true)
self.b -= lr * delta.sum()
self.w -= lr * torch.matmul(x.T, delta)gradient descent¶
@add_to_class(MultiLinearRegression)
def fit(self, x: torch.Tensor, y: torch.Tensor,
epochs: int, lr: float, batch_size: int,
x_valid: torch.Tensor, y_valid: torch.Tensor):
"""
Fit the model using gradient descent.
Args:
x: Input tensor of shape (n_samples, n_features).
y: Target tensor of shape (n_samples,).
epochs: Number of epochs to fit.
lr: learning rate.
batch_size: Int number of batch.
x_valid: Input tensor of shape (n_valid_samples, n_features).
y_valid: Target tensor of shape (n_valid_samples,).
"""
for epoch in range(epochs):
loss = []
for batch in range(0, len(y), batch_size):
end_batch = batch + batch_size
y_pred = self.predict(x[batch:end_batch])
loss.append(self.mse_loss(
y[batch:end_batch],
y_pred
))
self.update(
x[batch:end_batch],
y[batch:end_batch],
y_pred,
lr
)
loss = round(sum(loss) / len(loss), 4)
loss_v = round(self.evaluate(x_valid, y_valid), 4)
print(f'epoch: {epoch} - MSE: {loss} - MSE_v: {loss_v}')Scrath vs Torch.nn¶
Torch.nn model¶
class TorchLinearRegression(nn.Module):
def __init__(self, n_features):
super(TorchLinearRegression, self).__init__()
self.layer = nn.Linear(n_features, 1, device=device)
self.loss = nn.MSELoss()
def forward(self, x):
return self.layer(x)
def evaluate(self, x, y):
self.eval()
with torch.no_grad():
y_pred = self.forward(x)
return self.loss(y_pred, y).item()
def fit(self, x, y, epochs, lr, batch_size, x_valid, y_valid):
optimizer = torch.optim.SGD(self.parameters(), lr=lr)
for epoch in range(epochs):
loss_t = [] # train loss
for batch in range(0, len(y), batch_size):
end_batch = batch + batch_size
y_pred = self.forward(x[batch:end_batch])
loss = self.loss(y_pred, y[batch:end_batch])
loss_t.append(loss.item())
optimizer.zero_grad()
loss.backward()
optimizer.step()
loss_t = round(sum(loss_t) / len(loss_t), 4)
loss_v = round(self.evaluate(x_valid, y_valid), 4)
print(f'epoch: {epoch} - MSE: {loss_t} - MSE_v: {loss_v}')
optimizer.zero_grad()torch_model = TorchLinearRegression(N)scratch model¶
model = MultiLinearRegression(N)evals¶
We will use a metric to compare our model with the PyTorch model.
import MAPE modified¶
# This cell imports torch_mape
# if you are running this notebook locally
# or from Google Colab.
import os
import sys
module_path = os.path.abspath(os.path.join('..'))
if module_path not in sys.path:
sys.path.append(module_path)
try:
from tools.torch_metrics import torch_mape as mape
print('mape imported locally.')
except ModuleNotFoundError:
import subprocess
repo_url = 'https://raw.githubusercontent.com/PilotLeoYan/inside-deep-learning/main/content/tools/torch_metrics.py'
local_file = 'torch_metrics.py'
subprocess.run(['wget', repo_url, '-O', local_file], check=True)
try:
from torch_metrics import torch_mape as mape # type: ignore
print('mape imported from GitHub.')
except Exception as e:
print(e)mape imported locally.
predictions¶
Let’s compare the predictions of our model and PyTorch’s using modified MAPE.
mape(
model.predict(X_valid),
torch_model.forward(X_valid).squeeze(-1)
)13.10236558222539They differ considerably because each model has its own parameters initialized randomly and independently of the other model.
copy parameters¶
We copy the values of the PyTorch model parameters to our model.
model.copy_params(torch_model.layer)predictions after copy parameters¶
We measure the difference between the predictions of both models again.
mape(
model.predict(X_valid),
torch_model.forward(X_valid).squeeze(-1)
)0.0We can see that their predictions do not differ greatly.
loss¶
mape(
model.evaluate(X_valid, Y_valid),
torch_model.evaluate(X_valid, Y_valid.unsqueeze(-1))
)0.0training¶
We are going to train both models using the same hyperparameters’ value. If our model is well designed, then starting from the same parameters it should arrive at the same parameters’ values as the PyTorch model after training.
LR: float = 0.01 # learning rate
EPOCHS: int = 16 # number of epochs
BATCH: int = len(X_train) // 3 # number of minibatchtorch_model.fit(
X_train,
Y_train.unsqueeze(-1),
EPOCHS, LR, BATCH,
X_valid,
Y_valid.unsqueeze(-1)
)epoch: 0 - MSE: 11397.2866 - MSE_v: 8707.6512
epoch: 1 - MSE: 8993.6666 - MSE_v: 7320.302
epoch: 2 - MSE: 7155.589 - MSE_v: 6216.6786
epoch: 3 - MSE: 5744.2143 - MSE_v: 5330.7036
epoch: 4 - MSE: 4655.3057 - MSE_v: 4612.4923
epoch: 5 - MSE: 3810.5595 - MSE_v: 4024.314
epoch: 6 - MSE: 3151.1116 - MSE_v: 3537.5675
epoch: 7 - MSE: 2632.6743 - MSE_v: 3130.5135
epoch: 8 - MSE: 2221.892 - MSE_v: 2786.5756
epoch: 9 - MSE: 1893.6122 - MSE_v: 2493.0666
epoch: 10 - MSE: 1628.8397 - MSE_v: 2240.2322
epoch: 11 - MSE: 1413.2041 - MSE_v: 2020.5346
epoch: 12 - MSE: 1235.8103 - MSE_v: 1828.114
epoch: 13 - MSE: 1088.3777 - MSE_v: 1658.3839
epoch: 14 - MSE: 964.5936 - MSE_v: 1507.7271
epoch: 15 - MSE: 859.6285 - MSE_v: 1373.2668
model.fit(
X_train, Y_train,
EPOCHS, LR, BATCH,
X_valid, Y_valid
)epoch: 0 - MSE: 11397.2866 - MSE_v: 8707.6512
epoch: 1 - MSE: 8993.6666 - MSE_v: 7320.302
epoch: 2 - MSE: 7155.589 - MSE_v: 6216.6786
epoch: 3 - MSE: 5744.2143 - MSE_v: 5330.7036
epoch: 4 - MSE: 4655.3057 - MSE_v: 4612.4923
epoch: 5 - MSE: 3810.5595 - MSE_v: 4024.314
epoch: 6 - MSE: 3151.1116 - MSE_v: 3537.5675
epoch: 7 - MSE: 2632.6743 - MSE_v: 3130.5135
epoch: 8 - MSE: 2221.892 - MSE_v: 2786.5756
epoch: 9 - MSE: 1893.6122 - MSE_v: 2493.0666
epoch: 10 - MSE: 1628.8397 - MSE_v: 2240.2322
epoch: 11 - MSE: 1413.2041 - MSE_v: 2020.5346
epoch: 12 - MSE: 1235.8103 - MSE_v: 1828.114
epoch: 13 - MSE: 1088.3777 - MSE_v: 1658.3839
epoch: 14 - MSE: 964.5936 - MSE_v: 1507.7271
epoch: 15 - MSE: 859.6285 - MSE_v: 1373.2668
predictions after training¶
mape(
model.predict(X_valid),
torch_model.forward(X_valid).squeeze(-1)
)3.4679191706207704e-16bias¶
We directly measure the difference between the bias values of both models.
mape(
model.b.clone(),
torch_model.layer.bias.detach()
)0.0weight¶
And measure the difference between the weight values of both models.
mape(
model.w.clone(),
torch_model.layer.weight.detach().squeeze(0)
)7.449571250865014e-17All right, our implementation is correct respect to PyTorch. Now, we can finally tackle Multioutput in the next notebook.

